Date d'arrivée : 2003
Statut : Ingénieur d'étude - Directeur associé
Rôles :
Dernier diplôme obtenu : DESS architecture des systèmes d'information
Comprehensive annotation of olfactory and gustatory receptor genes and transposable elements revealed their evolutionary dynamics in aphids
Une erreur est survenue lors de la communication avec l'API PubMed.
Cliquez ici pour accéder à la publication sur le site de PubMed
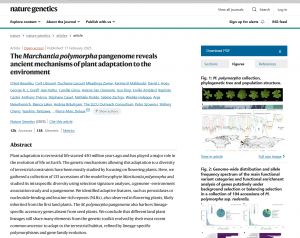
The Marchantia polymorpha pangenome reveals ancient mechanisms of plant adaptation to the environment
Une erreur est survenue lors de la communication avec l'API PubMed.
Cliquez ici pour accéder à la publication sur le site de PubMed
Genome assembly of a diversity panel of Chenopodium quinoa
Une erreur est survenue lors de la communication avec l'API PubMed.
Cliquez ici pour accéder à la publication sur le site de PubMed
An online database for einkorn wheat to aid in gene discovery and functional genomics studies
Une erreur est survenue lors de la communication avec l'API PubMed.
Cliquez ici pour accéder à la publication sur le site de PubMed
Einkorn genomics sheds light on history of the oldest domesticated wheat
Une erreur est survenue lors de la communication avec l'API PubMed.
Cliquez ici pour accéder à la publication sur le site de PubMed
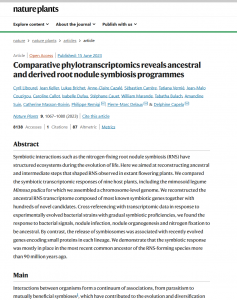
Comparative phylotranscriptomics reveals ancestral and derived root nodule symbiosis programmes
Une erreur est survenue lors de la communication avec l'API PubMed.
Cliquez ici pour accéder à la publication sur le site de PubMed
The genomics of linkage drag in inbred lines of sunflower
Une erreur est survenue lors de la communication avec l'API PubMed.
Cliquez ici pour accéder à la publication sur le site de PubMed
Long-read genome sequencing of bread wheat facilitates disease resistance gene cloning
Une erreur est survenue lors de la communication avec l'API PubMed.
Cliquez ici pour accéder à la publication sur le site de PubMed
A genome sequence resource for the genus Passiflora, the genome of the wild diploid species Passiflora organensis
Une erreur est survenue lors de la communication avec l'API PubMed.
Cliquez ici pour accéder à la publication sur le site de PubMed
Population genomics of apricots unravels domestication history and adaptive events
Une erreur est survenue lors de la communication avec l'API PubMed.
Cliquez ici pour accéder à la publication sur le site de PubMed
Fonio millet genome unlocks African orphan crop diversity for agriculture in a changing climate
Une erreur est survenue lors de la communication avec l'API PubMed.
Cliquez ici pour accéder à la publication sur le site de PubMed
A repertory of rearrangements and the loss of an inverted repeat region in Passiflora chloroplast genomes
Une erreur est survenue lors de la communication avec l'API PubMed.
Cliquez ici pour accéder à la publication sur le site de PubMed
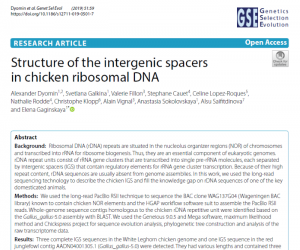
Structure of the intergenic spacers in chicken ribosomal DNA
Une erreur est survenue lors de la communication avec l'API PubMed.
Cliquez ici pour accéder à la publication sur le site de PubMed
A receptor-like kinase enhances sunflower resistance to Orobanche cumana
Une erreur est survenue lors de la communication avec l'API PubMed.
Cliquez ici pour accéder à la publication sur le site de PubMed
Influence of CNV on transcript levels of HvCBF genes at Fr-H2 locus revealed by resequencing in resistant barley cv. ‘Nure’ and expression analysis
Une erreur est survenue lors de la communication avec l'API PubMed.
Cliquez ici pour accéder à la publication sur le site de PubMed
μLAS technology for DNA isolation coupled to Cas9-assisted targeting for sequencing and assembly of a 30 kb region in plant genome
Une erreur est survenue lors de la communication avec l'API PubMed.
Cliquez ici pour accéder à la publication sur le site de PubMed
A gene-rich fraction analysis of the Passiflora edulis genome reveals highly conserved microsyntenic regions with two related Malpighiales species
Une erreur est survenue lors de la communication avec l'API PubMed.
Cliquez ici pour accéder à la publication sur le site de PubMed
Whole-genome landscape of Medicago truncatula symbiotic genes
Une erreur est survenue lors de la communication avec l'API PubMed.
Cliquez ici pour accéder à la publication sur le site de PubMed
Genome-wide identification of the mutation underlying fleece variation and discriminating ancestral hairy species from modern woolly sheep.
Une erreur est survenue lors de la communication avec l'API PubMed.
Cliquez ici pour accéder à la publication sur le site de PubMed
Genome-wide identification of the mutation underlying fleece variation and discriminating ancestral hairy species from modern woolly sheep.
Demars J, Cano M, Drouilhet L, Plisson-Petit F, Bardou P, Fabre S, Servin B, Sarry J, Woloszyn F, Mulsant P, Foulquier D, Carriere F, Aletru M, Rodde N, Cauet S, Bouchez O, Pirson M, Tosser-Klopp G, Allain D.
Journal: Molecular Biology and Evolution
DOI: 34: 1722-1729.
Voir plus
Demars J, Cano M, Drouilhet L, Plisson-Petit F, Bardou P, Fabre S, Servin B, Sarry J, Woloszyn F, Mulsant P, Foulquier D, Carriere F, Aletru M, Rodde N, Cauet S, Bouchez O, Pirson M, Tosser-Klopp G, Allain D.
Journal: Molecular Biology and Evolution
DOI: 34: 1722-1729.
Abstract
The composition and structure of fleece variation observed in mammals is a consequence of a strong selective pressure for fiber production after domestication. In sheep, fleece variation discriminates ancestral species carrying a long and hairy fleece from modern domestic sheep (Ovis aries) owning a short and woolly fleece. Here, we report that the “woolly” allele results from the insertion of an antisense EIF2S2 retrogene (called asEIF2S2) into the 3′ UTR of the IRF2BP2 gene leading to an abnormal IRF2BP2 transcript. We provide evidence that this chimeric IRF2BP2/asEIF2S2 messenger 1) targets the genuine sense EIF2S2 RNA and 2) creates a long endogenous double-stranded RNA which alters the expression of both EIF2S2 and IRF2BP2 mRNA. This represents a unique example of a phenotype arising via a RNA-RNA hybrid, itself generated through a retroposition mechanism. Our results bring new insights on the sheep population history thanks to the identification of the molecular origin of an evolutionary phenotypic variation.
Link: https://academic.oup.com/mbe/article/34/7/1722/3091105
The chloroplast genome of Passiflora edulis (Passifloraceae) assembled from long sequence reads: structural organization and phylogenomic studies in Malpighiales
Une erreur est survenue lors de la communication avec l'API PubMed.
Cliquez ici pour accéder à la publication sur le site de PubMed
The chloroplast genome of Passiflora edulis (Passifloraceae) assembled from long sequence reads: structural organization and phylogenomic studies in Malpighiales
Front. Plant Sci. 8:334. DOI: 10.3389/fpls.2017.00334
Voir plus
Authors :
Cauz Santos LA, Freitas Munhoz C, Rodde N, Cauet S, Azevedo Santos A, Alves Penha H, Carnier Dornelas M, de Mello Varani A, Conde Xavier Oliveira G, Bergès H, Carneiro Vieira ML (2017)
Front. Plant Sci. 8:334. DOI: 10.3389/fpls.2017.00334
Abstract :
The family Passifloraceae consists of some 700 species classified in around 16 genera. Almost all its members belong to the genus Passiflora. In Brazil, the yellow passion fruit (Passiflora edulis) is of considerable economic importance, both for juice production and consumption as fresh fruit. The availability of chloroplast genomes (cp genomes) and their sequence comparisons has led to a better understanding of the evolutionary relationships within plant taxa. In this study, we obtained the complete nucleotide sequence of the P. edulis chloroplast genome, the first entirely sequenced in the Passifloraceae family. We determined its structure and organization, and also performed phylogenomic studies on the order Malpighiales and the Fabids clade. The P. edulis chloroplast genome is characterized by the presence of two copies of an inverted repeat sequence (IRA and IRB) of 26,154 bp, each separating a small single copy region of 13,378 bp and a large single copy (LSC) region of 85,720 bp. The annotation resulted in the identification of 105 unique genes, including 30 tRNAs, 4 rRNAs, and 71 protein coding genes. Also, 36 repetitive elements and 85 SSRs (microsatellites) were identified. The structure of the complete cp genome of P. edulis differs from that of other species because of rearrangement events detected by means of a comparison based on 22 members of the Malpighiales. The rearrangements were three inversions of 46,151, 3,765 and 1,631 bp, located in the LSC region. Phylogenomic analysis resulted in strongly supported trees, but this could also be a consequence of the limited taxonomic sampling used. Our results have provided a better understanding of the evolutionary relationships in the Malpighiales and the Fabids, confirming the potential of complete chloroplast genome sequences in inferring evolutionary relationships and the utility of long sequence reads for generating very accurate biological information.
Long Read Sequencing Technology to Solve Complex Genomic Regions Assembly in Plants.
Une erreur est survenue lors de la communication avec l'API PubMed.
Cliquez ici pour accéder à la publication sur le site de PubMed
Long Read Sequencing Technology to Solve Complex Genomic Regions Assembly in Plants.
Journal of Next Generation Sequencing & Applications.
Voir plus
Authors :
Arnaud Bellec, Audrey Courtial, Stephane Cauet, Nathalie Rodde, Sonia Vautrin, Genseric Beydon, Nadege Arnal, Nadine Gautier, Joelle Fourment, Elisa Prat, William Marande, Yves Barriere and Helene Berges.
Journal of Next Generation Sequencing & Applications.
Abstract :
Background:
Numerous completed or on-going whole genome sequencing projects have highlighted the fact that obtaining a high quality genome sequence is necessary to address comparative genomics questions such as structural variations among genotypes and gain or loss of specific function. Despite the spectacular progress that has been made in sequencing technologies, obtaining accurate and reliable data is still a challenge, both at the whole genome scale and when targeting specific genomic regions. These problems are even more noticeable for complex plant genomes. Most plant genomes are known to be particularly challenging due to their size, high density of repetitive elements and various levels of ploidy. To overcome these problems, we have developed a strategy to reduce genome complexity by using the large insert BAC libraries combined with next generation sequencing technologies.
Results:
We compared two different technologies (Roche-454 and Pacific Biosciences PacBio RS II) to sequence pools of BAC clones in order to obtain the best quality sequence. We targeted nine BAC clones from different species (maize, wheat, strawberry, barley, sugarcane and sunflower) known to be complex in terms of sequence assembly. We sequenced the pools of the nine BAC clones with both technologies. We compared assembly results and highlighted differences due to the sequencing technologies used.
Conclusions:
We demonstrated that the long reads obtained with the PacBio RS II technology serve to obtain a better and more reliable assembly, notably by preventing errors due to duplicated or repetitive sequences in the same region.
Link :
Une erreur est survenue lors de la communication avec l'API PubMed.
Cliquez ici pour accéder à la publication sur le site de PubMed