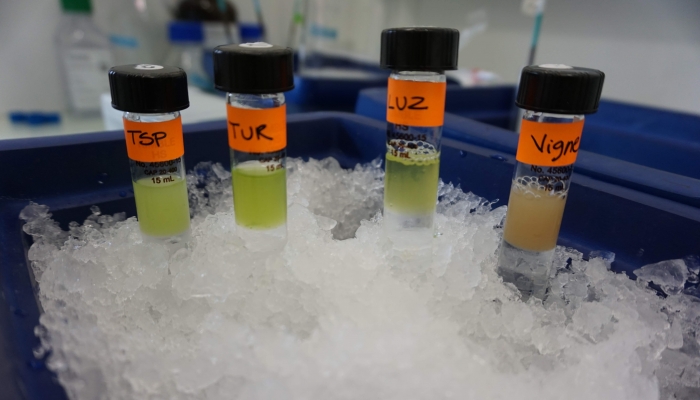
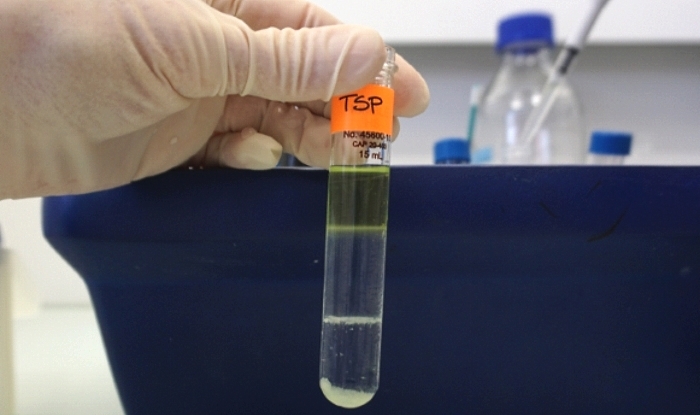
Genome optical mapping

Optical mapping is a technique that relies on the Saphyr system from Bionano to obtain the physical map, so called Optical Map, of a genome.
|
| The CNRGV is a certified service provider approved by Bionano for plant and animals genomes optical mapping. This label recognizes the conformity of our process with guidelines and our expertise to providethe best possible data. |
Based on in-silico assembly of large DNA molecules labelled at specific sites, the Optical Map provides long-distance information revealing the structure of studied genome.
The Optical Map is then combined with sequence information to provide a high quality scaffolded genome assembly called hybrid assembly.
The CNRGV offer to build the Optical Map of your genome of interest from the extraction of the DNA to the combination of the data generated by the Saphyr system with your sequencing data.
Service workflow:
- Isolation of High Molecular Weight DNA from plant tissues
- Direct labeling of the DNA (DLE production)
- Loading prepared DNA on a Saphyr Chip
- Imaging of linearized DNA molecules by the Saphyr system
- Data analysis using Access software to build the optical map of the genome
Optical Mapping with BioNano Technology General workflow
Optical maps are powerful tools for understanding the structures and functions of genomes. They are particularly useful for scaffolding de novo sequences and for detecting structural variations and repetitions within complex genomes.
Comparing several individual optical maps will allow you to highlight the structural variations and sequence copy number variation while providing positional information.
Example of Optical maps to improve the assembly quality of genome:
|
| Sunflower
|
|
| Polyploid wheat
The longest hybrid scaffold was 627.2 Mb and covered 99% of chromosome 3D. Publication: Long-read genome sequencing of bread wheat facilitates disease resistance gene cloning
|
|
| Apricot tree
In this project, this apricot genome was the reference to be compared with three other genomes, produced using the same assembly strategy at various levels.We used OMAP to produce a catalog of large SV (insertion/deletion, duplication and inversion) for the 3 genomes compared to the reference. This approach is complementary to genome comparison through sequences alignments, bringing validation of SV detected through contigs comparison. Publication: Population genomics of apricots unravels domestication history and adaptive events |

 Genomic samples storage & distribution
Genomic samples storage & distribution Large DNA Fragment cloning and characterization
Large DNA Fragment cloning and characterization HMW DNA extraction
HMW DNA extraction Genome optical mapping
Genome optical mapping Genome scaffolding using Hi-C / Omni-C
Genome scaffolding using Hi-C / Omni-C Whole genome characterization
Whole genome characterization Large DNA Fragment Capture
Large DNA Fragment Capture









